Finding Correlations in Functionally Equivalent Proteins
Learning about the construction rules for proteins
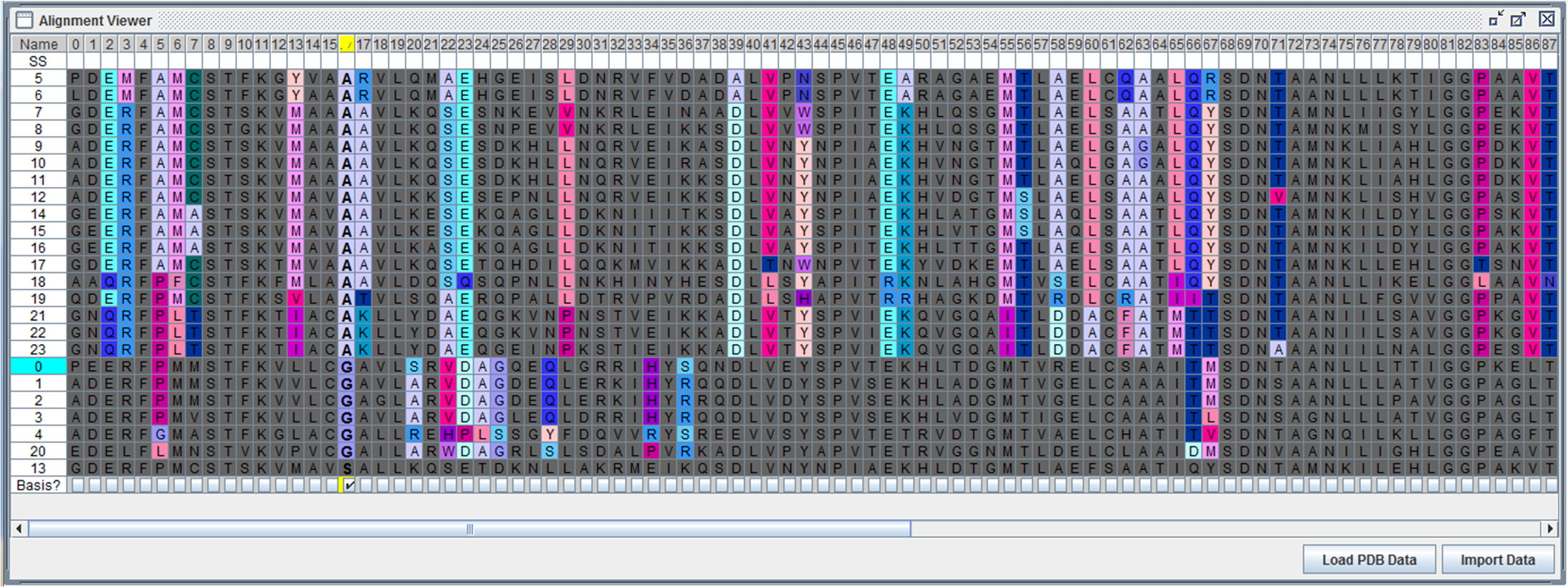
In recent years the amount of protein sequence data has grown explosively. Much interesting information is hidden in the databases and possibly could answer many questions of scientific and medicinal interest. This project addresses the question what restrictions are protein sequences subjected to.
Using alignments of functionally equivalent proteins, regularities such as correlated positions or residue patterns can be searched for. These regularities are assumed to be necessary to ensure a specific fold and various cellular functions. The developed visual analytics tool VisAlign supports the analysis by combining an automatic calculation of correlations with an interactive visualization.

